32106 FnCas12a (Cpf1) Nuclease
32106 FnCas12a (Cpf1) Nuclease
-
Specifications:
- 100 pmoL
- 1,000 pmoL
Article number:
32106
Price:Contact our distributors
隐藏域元素占位
Product overview
Description
FnCas12a (previously known as FnCpf1), a DNA endonuclease guided by crRNA alone, is derived from the Francisella tularensis strain. FnCas12a can specifically cleave the dsDNA target in a PAM (TTN)-dependent manner, generating DSBs, and cleave ssDNA target in a PAM-independent manner. In addition, both dsDNA and ssDNA targets can trigger the trans-cleavage activity of FnCas12a, that is, when Cas12a binds crRNA and target DNA to form a ternary complex, the trans-cleavage activity of Cas12a will be released to non-specifically cleave ssDNA sequences in the reaction system. Therefore, FnCas12a can be used not only for specific cleavage of dsDNA in vitro, but also for rapid detection of target nucleic acid. For details, please refer to the HOLMES rapid nucleic acid detection technology developed by Tolo Biotech (for details, please refer to PMID: 29707234).
Product Components
| Components |
32106-01 (100 pmol) |
32106-03 (1,000 pmol) |
|
|
100 pmol |
1,000 pmol |
|
|
1 mL |
1 mL |
a.The concentration of FnCas12a Nuclease is 10 μM (10 pmol/μL);
Storage Conditions
Store at -20℃; transport at ≤0℃.
Experimental Results
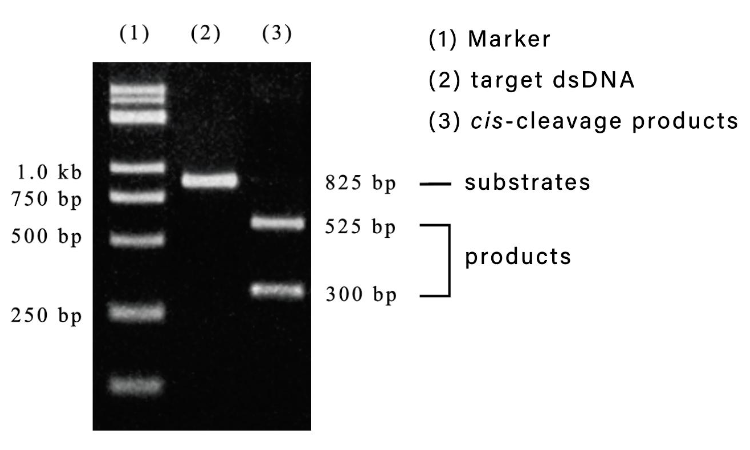
Fig1:Cis-cleavage

Fig2:Trans-cleavage
Experiment Procedure
1.Cis-cleavage
|
Components |
Volume |
Final concentration |
|
10 × HOLMES Buffer 1 |
2 μL | 1 × |
|
10 μM FnCas12a Nuclease |
0.5 μL | 250 nM |
|
10 μM crRNA |
0.5 μL | 250 nM |
|
1 μM Target DNA |
0.5 μL | 25 nM |
|
Nuclease-free water |
Up to 20 μL |
◆ Target DNA can be ssDNA or dsDNA with PAM sequence in the cis-cleavage system.
◆ If nucleic acid electrophoresis is used to analyze cis-cleavage products, it is recommended to use dsDNA targets because ssDNA targets will be further cleaved by trans-cleavage-activated Cas12a. For a 20 μL cis-cleavage reaction system, the recommended amount of target DNA is 100 ng to 500 ng, and the length of the target DNA fragment is in the range of 300 bp to 3 kb. The amount of Cas enzyme and crRNA required for different lengths of target DNA needs to be calculated according to the molar amount of target DNA. It is recommended to keep the molar ratio of Cas enzyme:crRNA:target DNA at 10:10:1 to ensure that the target DNA is cleaved completely.
◆ Incubate at 37°C for 30 min~1 h, inactivate at 85°C for 5 min, then analyze the cis-cleaved products by nucleic acid electrophoresis.
2. Trans-cleavage experiment
|
Components |
Volume |
Final concentration |
|
10 × HOLMES Buffer 1 |
2 μL | 1 × |
|
10 μM FnCas12a Nuclease |
0.05~0.5 μL | 25~250 nM |
|
10 μM crRNA |
0.05~0.5 μL | 25~250 nM |
|
1 μM Target DNA |
0.5~5 μL | 25~250 nM |
|
10 μM ssDNA Reporter |
0.05~0.5 μL | 25~250 nM |
|
Nuclease-free water |
Up to 20 μL |
◆ Target DNA can be ssDNA or dsDNA with PAM sequence in the trans-cleavage system.
◆ The amount of each component in the trans-cleavage system can be adjusted according to different experimental purposes. When the amount is small, each component can be diluted first and then added to the system. FnCas12a Nuclease can be diluted with 1×HOLMES Buffer 1, and it must be used immediately after dilution (If you want to store the diluted Cas12 enzyme for a long term, please use Cas12 Dilution Buffer, #32012). crRNA, Target DNA and ssDNA Reporter can be diluted with Nuclease-free Water, but for extremely low concentrations of Target DNA (such as LOD experiments), it is recommended to dilute with 0.1 % Tween 20 and use low adsorption centrifuge tubes and tips etc.
◆ For the amount of probe used in the trans-cleavage system and the expected reaction time to reach the plateau, please refer to the specific value of transU of each batch of Cas enzyme. For details, please refer to the COA test report provided by Tolo Biotech.
◆ The real-time fluorescent quantitative PCR instrument detects the fluorescent signal from reaction at 37°C, and collects the fluorescent signal every 30 seconds.
FAQ
Q1: What are the differences between Cas12a and Cas9?
A1: In general, Cas9 requires the guidance of crRNA and tracrRNA together (or a fused sgRNA) to specifically recognize and cleave dsDNA targets with a PAM (5´-NGG-3´) sequence and does not possess trans-cleavage activity. On the other hand, Cas12a only requires crRNA for guidance, can specifically recognize and cleave dsDNA targets with a PAM (5´TTTN-3´ or 5´TTN-3´) sequence, and does not rely on a PAM sequence for the recognition and cleavage of ssDNA targets. Additionally, both dsDNA and ssDNA targets can activate Cas12a's trans-cleavage activity, allowing it to non-specifically cleave ssDNA in the system. A detailed comparison is provided in the table below:
|
Cas9 |
Cas12a (Cpf1) |
|
|
Classification |
Class II Type II |
Class II Type V |
|
Nuclease Domains |
RuvC and HNH |
RuvC only |
|
Guide RNA |
crRNA and tracrRNA |
crRNA only |
|
PAM Sequence |
5´-NGG-3´ |
5´TTTN-3´ or 5´TTN-3´ |
|
Cis-Cleavage Target |
dsDNA with PAM |
dsDNA with PAM or ssDNA |
|
Cis-Cleavage Site |
3 bases 5´ of the PAM |
~18 bases 3´ of the PAM |
|
Termini of Cis-Cleavage Products |
Blunt ends |
5´ overhang |
|
Trans-Cleavage Target |
/ |
ssDNA |
Q2: How to design the crRNA for FnCas12a, and does the length of the crRNA impact FnCas12a's cleavage efficiency?
A2:The crRNA for FnCas12a consists of a 19-nt full-length direct repeat sequence (DR region, also known as the scaffold region) and a 16-30 nt guide sequence (spacer region). For dsDNA targets, the length of the crRNA guide sequence should be at least 16 nt, as guide sequences of 14 or 15 nt exhibit weak cleavage activity. For ssDNA targets, the guide sequence length can even be truncated to 10 nt while still retaining cleavage activity. The crRNA for FnCas12a can be designed based on the following sequence:
5′– AAUUUCUACUGUUGUAGAUNNNNNNNNNNNNNNNNNNNN–3′, (The underlined portion represents the spacer region that complements the specific target sequence).
Q3: Do Cas12a enzymes from different sources recognize different PAM sequences?
A3: Cas12a enzyme family typically recognizes PAM sequences rich in thymidine (T). For example, FnCas12a and MbCas12a recognize 5´-TTN-3´ PAM sequences, while AsCas12a, LbCas12a and Lb2Cas12a recognize 5´-TTTN-3´ PAM sequences.
Q4: Besides FnCas12a, what are some other common Cas12a enzymes, and how do their direct repeat sequences (also known as the DR region or scaffold region) differ?
A4: Common Cas12a enzymes include FnCas12a, LbCas12a, AsCas12a, etc. The direct repeat sequences (DR regions or scaffold regions) of crRNAs within the Cas12a family are highly conserved. Below is a table showing the conservation of the DR region sequences for various Cas12a enzymes (please note that the first base at the 5' end of the crRNA is typically processed by Cas12a, so it may be omitted when designing crRNAs):

Q5: How should the ratio of Cas12a enzyme: crRNA: target DNA be adjusted in the cis-cleavage system?
A5: To ensure complete cleavage of the target dsDNA, it is conventionally recommended to maintain a Cas12a enzyme: crRNA: target DNA ratio of 10:10:1. If crRNA efficiency is low, it is suggested to try a Cas12a enzyme: crRNA ratio of 1:1.25~2. Excessive amounts of crRNA can also negatively affect Cas12a cleavage efficiency. Additionally, if the crRNA is prepared by transcription, it is important to ensure the purity of the transcription product.
Q6: Can both dsDNA targets and ssDNA targets activate Cas12a's trans-cleavage activity, and which is more efficient?
A6: Both dsDNA and ssDNA targets can activate Cas12a's trans-cleavage activity (also known as collateral cleavage), similar to Cas12b. However, ssDNA targets typically stimulate the trans-activity of Cas12b more efficiently, while dsDNA targets usually activate the trans-activity of Cas12a more effectively.
Q7: What determines the cleavage sites for Cas12a on dsDNA targets and ssDNA targets, and where specifically does this occur?
A7: The cleavage sites for Cas12a on dsDNA targets primarily depend on the presence of a PAM sequence. Previous research has shown that Cas12a cleaves the non-target strand (NTS) around position 18 from the PAM and the target strand (TS) around position 23 from the PAM, resulting in a 5' overhang of 5 nucleotides. However, these cleavage sites are not highly specific and can vary depending on the source of Cas12a, the target sequence, and the length of the crRNA. In particular, cleavage on the NTS can occur around positions 14 (12th-15th) and 18 (18th-20th) from the PAM, while cleavage on the TS can occur around position 22 (21st-24th) from the PAM. Cas12a enzyme family does not rely on a PAM sequence for cleaving ssDNA targets, and the cleavage site is typically located around position 22 (21st-23th) from the first 3'-base that pairs with the crRNA guide sequence.
Q8: Is the occurrence of cis-cleavage a prerequisite for FnCas12a to activate and non-specifically trans-cleave ssDNA probes?
A8: No, ss DNA targets of length 18 nt that do not contain the cis-cleavage site (21st-23th) can also activate Cas12a's trans-cleavage activity. This suggests that the occurrence of cis-cleavage is not a prerequisite for trans-cleavage.
References
Related Products






 FnCas12a Nuclease (10 μM) a
FnCas12a Nuclease (10 μM) a 10 × HOLMES Buffer 1
10 × HOLMES Buffer 1



















