32101 SpCas9 Nuclease
32101 SpCas9 Nuclease
-
Specifications:
- 100 pmoL
- 1,000 pmoL
Article number:
32101
Price:Contact our distributors
隐藏域元素占位
Product overview
Description
SpCas9 is a DNA endonuclease guided by crRNA and tracrRNA (or sgRNA alone), which is derived from S. pyogenes. SpCas9 specifically recognizes and cleaves dsDNA target in a PAM (NGG)-dependent manner. The cleavage site of Cas9 in the target sequence is 3 bps away from the PAM site. In addition, SpCas9 can also specifically cleave ssDNA or ssRNA in the presence of DNA PAMmer sequences (for details, please refer to PMID: 24476820; PMID: 25274302). SpCas9 can also be used in in vitro experiments for cleaving target DNA and cloning target fragments (for details, please refer to PMID: 26323354).
Product Components
|
Components |
32101-01(100 pmol) |
32101-03(1,000 pmol) |
|
|
100 pmol | 1,000 pmol |
|
|
1 mL | 1 mL |
a.The concentration of SpCas9 Nuclease is 10 μM (10 pmol/μL);
Storage Conditions
Store at -20℃; transport at ≤0℃.
Experimental Results
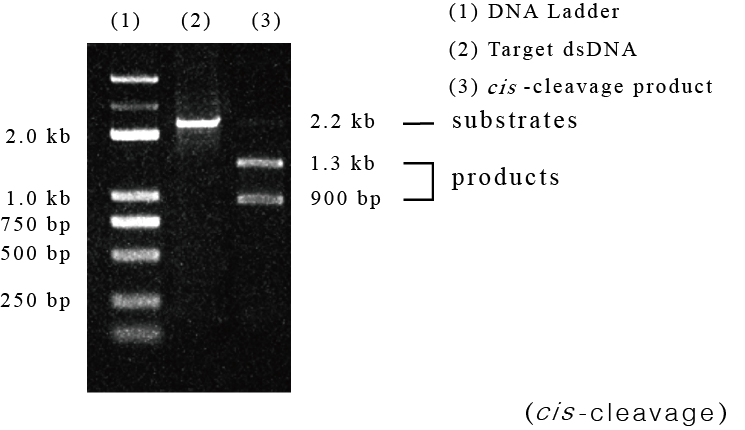
In this experiment, the dsDNA target was a 2.2-kb fragment of DNMT1 gene. The sgRNA designed in this experiment contained a spacer region complementary to the DNMT1 sequence. The experimental results showed that SpCas9 specifically recognized and cleaved the DNMT1 sequence under the guidance of sgRNA, generating two cis-cleaved products of 1.3 kb and 0.9 kb, respectively.
Fig1. SpCas9 specifically recognizes and cleaves target dsDNA under the guidance of sgRNA.
Experiment Procedure
1.Cis-cleavage
|
Components |
Volume |
Final concentration |
|
10 × TOLO Buffer 3 |
2 μL | 1 × |
|
10 μM SpCas9 Nuclease |
0.5 μL | 250 nM |
|
10 μM sgRNA |
0.5 μL | 250 nM |
|
1 μM Target dsDNA |
0.5 μL | 25 nM |
|
Nuclease-free water |
Up to 20 μL |
◆ If nucleic acid electrophoresis is used to analyze cis-cleavage products, for a 20 μL cis-cleavage reaction system, the recommended amount of target DNA is 100 ng to 500 ng, and the length of the target DNA fragment is in the range of 300 bp to 3 kb. The amount of Cas enzyme and sgRNA required for different lengths of target DNA needs to be calculated according to the molar amount of target DNA. It is recommended to keep the molar ratio of Cas enzyme:sgRNA:target DNA at 10:10:1 to ensure that the target DNA is cleaved completely.
◆ Incubate at 37°C for 30 min~1 h, inactivate at 85°C for 5 min, then analyze the cis-cleaved products by nucleic acid electrophoresis.
FAQ
Q1: What are the differences between Cas9, Cas12, and Cas13?
A1: The main difference between Cas9, Cas12 and Cas13 is that Cas12, when bound to guide RNA and target DNA, is activated for trans-cleavage activity for single-stranded DNA (ssDNA), whereas Cas13, when bound to guide RNA and target RNA, is activated for trans-cleavage activity for single-stranded RNA (ssRNA). Cas9, on the other hand, has not been reported to have trans-cleavage activity. A detailed comparison is provided in the table below:
|
|
Cas9 |
Cas12 |
Cas13 |
|
Classification |
Class II Type II |
Class II Type V |
Class II Type VI |
|
Nuclease Domains |
RuvC and HNH |
RuvC only |
HEPN × 2 |
|
Guide RNA |
crRNA and tracrRNA |
crRNA only or crRNA and tracrRNA |
crRNA only |
|
PAM Sequence |
5´-NGG-3´ |
T-Rich |
/ |
|
Cis-Cleavage Target |
dsDNA with PAM |
dsDNA with PAM or ssDNA |
ssRNA |
|
Trans-Cleavage Target |
/ |
ssDNA |
ssRNA |
Q2: How to design the sgRNA for SpCas9?
A2: The sgRNA for SpCas9 can be designed based on the following sequence:
5′–NNNNNNNNNNNNNNNNNNNNGUUUUAGAGCUAUGCUGAAAGCAUAGCAAGUUAAAAUAAGGCUAGUCCGUUAUCAACUUGAAAAAGUGGCACCGAGUCGGUG–3′, (The underlined portion represents the spacer region that complements the specific target sequence).
Q3: Are there any structural characteristics of the Cas9 domains responsible for cleaving dsDNA targets?
A3: Cas9 is a dual RNA-guided endonuclease that requires both crRNA and tracrRNA (or engineered fused sgRNA) for the recognition and cleavage of the target dsDNA. The mechanism involves two structural domains, RuvC and HNH. RuvC cleaves the non-target strand (NTS), while HNH cleaves the target strand (TS). Together, they create double-strand DNA breaks (DSBs) forming blunt ends, with the cleavage site located 3 base pairs from the PAM 5´-end in the target dsDNA sequence.
Q4: Can Cas9 only recognize and cleave dsDNA targets with a PAM?
A4: No, the Cas9 family is divided into subtypes A, B, and C, with C-type Cas9s shown to have a preference for cleaving ssDNA targets. Additionally, in the presence of a PAMmer, Cas9 can also cleave ssRNA targets.
Q5: Why is it recommended to maintain a molar ratio of 10:10:1 for the cis-cleavage system of Cas9 enzyme:sgRNA:target DNA?
A5: To ensure complete cleavage of the target dsDNA, it is conventionally recommended to maintain a Cas9 enzyme:sgRNA:target DNA ratio of 10:10:1. If sgRNA efficiency is low, it is suggested to try a Cas9 enzyme:sgRNA ratio of 1:1.25~2.
References
Related Products






 SpCas9 Nuclease (10 μM) a
SpCas9 Nuclease (10 μM) a 10 × TOLO Buffer 3
10 × TOLO Buffer 3